Tuan Le

I work on generative models for molecular design at Pfizer. My work sits at the intersection of probability, statistics, geometry, and chemistry. I’m interested in building systems that can generate molecules across several settings, including small-molecule drug discovery, protein sequence design, and other inverse problems in molecular design.
I earned a Ph.D. in computer science from Freie Universität Berlin, where I worked with Frank Noé on machine learning for molecular design. My Ph.D. research also grew out of a close collaboration with Djork-Arné Clevert. I’ve continued that collaboration at Bayer and Pfizer, working with teams in computational chemistry and machine learning.
Research interests
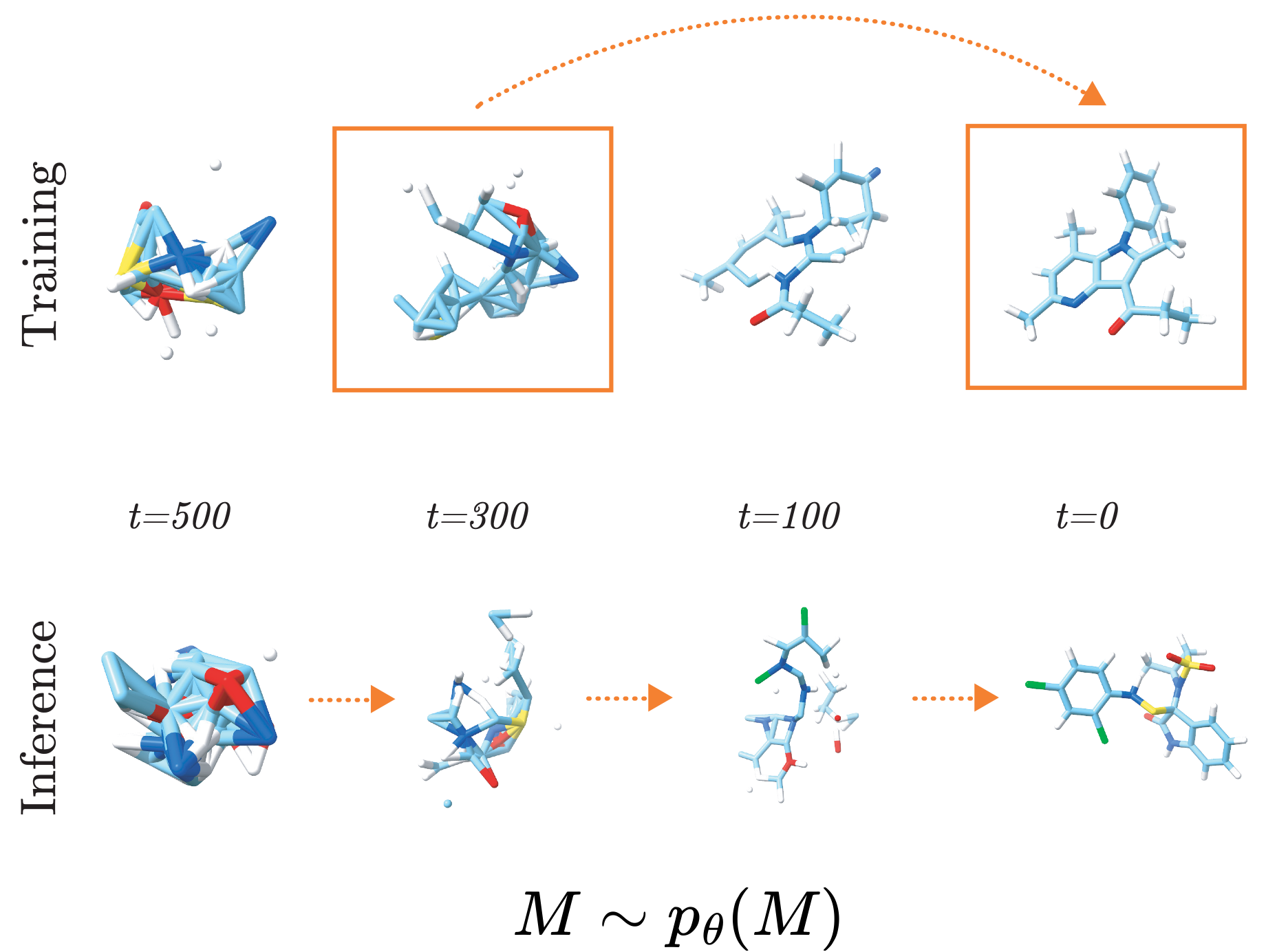
I’m interested in how generative models, especially diffusion models and flow matching, can be used for inverse design across molecular domains. That includes small-molecule drug discovery, where the goal may be to generate candidates for a target or property, as well as protein and sequence design.
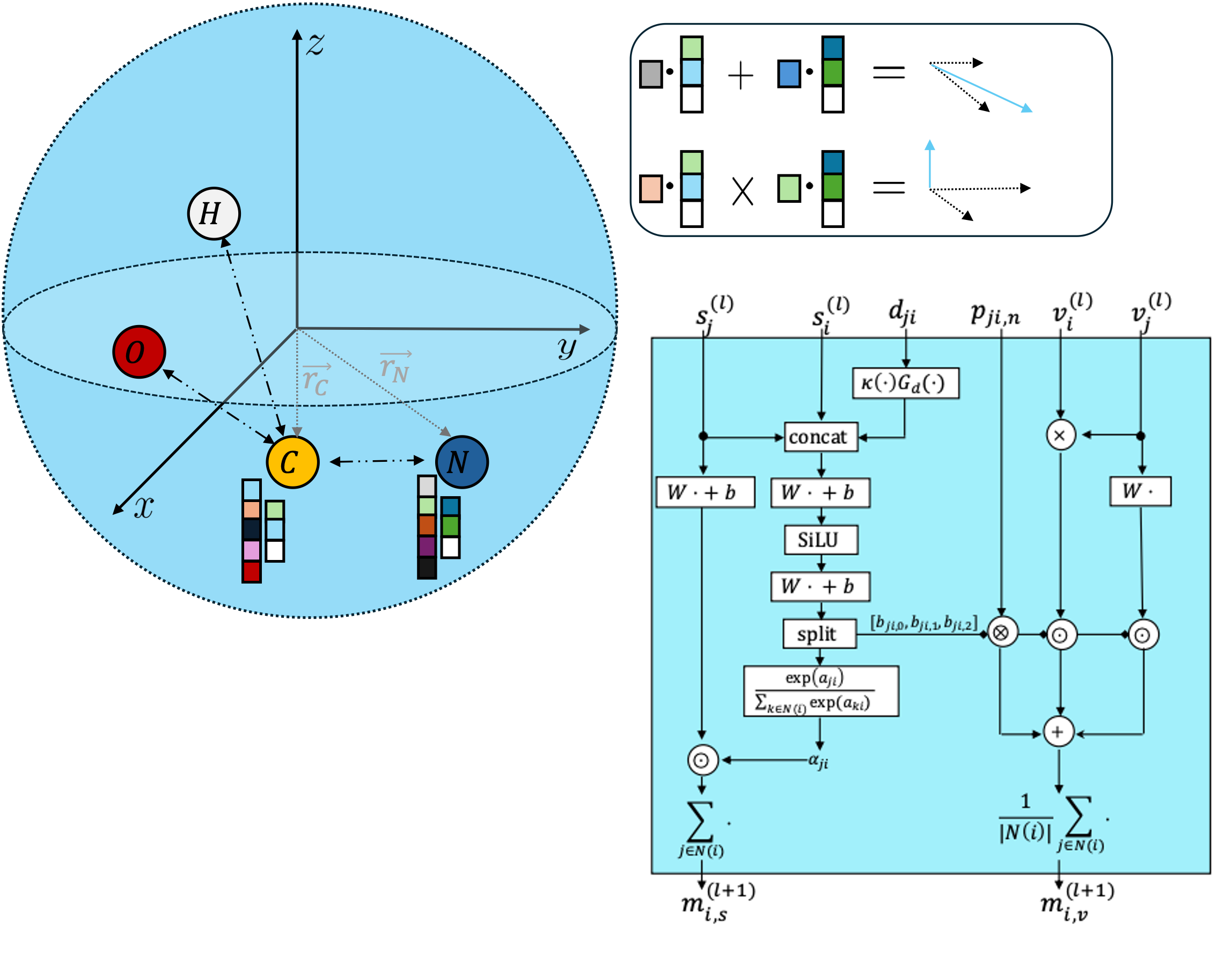
I also care about how domain knowledge enters the model. In my experience, the math, physics, and chemistry behind the problem should shape the system itself. Respecting physical structure, using 3D geometry, conditioning on the right information, and building in domain-specific features usually leads to more realistic and more useful molecules.
Just as important to me is the step after the method paper: turning a model into something people can actually use. Strong benchmark results matter, but they are only part of the story. The work becomes more useful when it fits the way chemists and other domain experts actually work.